Fast Kalman filtering and forward-backward smoothing via a
low-rank perturbative approach
Eftychios Pnevmatikakis, Kamiar Rahnama Rad,
Jonathan Huggins, and Liam Paninski
Under review
Kalman filtering-smoothing is a fundamental tool in statistical time
series analysis: it implements the optimal Bayesian filter in the
linear-Gaussian setting, and serves as a key step in the inference
algorithms for a wide variety of nonlinear and non-Gaussian
models. However, standard implementations of the Kalman filter require
$O(d^3)$ time and $O(d^2)$ space per timestep, where $d$ is the
dimension of the state variable, and are therefore impractical in
high-dimensional problems. In this paper we note that if a relatively
small number of observations are available per time step, the Kalman
equations may be approximated in terms of a low-rank perturbation of
the prior state covariance matrix in the absence of any observations.
In many cases this approximation may be computed and updated very
efficiently (often in just $O(k^2d)$ or $O(k^2d+kd\log d)$ time and
space per timestep, where $k$ is the rank of the perturbation and in
general $k\ll d$), using fast methods from numerical linear
algebra. We justify our approach and give bounds on the rank of the
perturbation as a function of the desired accuracy. For the case of
smoothing we also quantify the error of our algorithm due to the low
rank approximation and show that it can be made arbitrarily low at the
expense of a moderate computational cost. We describe applications
involving smoothing of spatiotemporal neuroscience data.
Preprint here;
see also:
- Paninski, L. (2010), Fast Kalman filtering on quasilinear
dendritic trees, Journal of Computational Neuroscience 28: 211-28;
- Pnevmatikakis, E. and Paninski, L. (2012), Fast
interior-point inference in high-dimensional sparse, penalized
state-space models, AISTATS '12.
Here are a couple examples. 
In this first example above, the black
dot represents the two-dimensional position x(t) of a mouse wandering
around its cage. The position of the mouse was recorded
simultaneously with the activity of neurons in the hippocampal
formation by Pablo Jercog, who graciously shared this data with us.
See Kentros et
al 2004 or Muzzio
et al 2009 for further details on experimental methods. We
applied our filtering methods to estimate the place field f[x,t] of a
single neuron as a function of two-dimensional position x and time t:
as in the simulated one-dimensional example discussed in the paper,
f[x,t] represents the expected firing rate as a function of position x
and time t, and is therefore explicitly allowed to change as a
function of time. The neuron's observed output at time t, y(t), is
represented as y(t)=f[x(t),t]+e(t), where e(t) is a random noise
source. The top panels display the fast Kalman filter-smoother
output at time t given the observations y(1),...,y(T) at the positions
x(1),...,x(T), where T is the length of the experiment (about 500
seconds here), while the bottom panels display the posterior marginal
variance (the posterior uncertainty about f[x,t]).
In more detail, we downsampled the position and neural firing rate
data to .66 Hz. The spike count in each timebin was transformed to
obtain the observation y(t); we applied a standard Anscombe
transformation to stabilize the variance of y(t). Under the
assumption that the spike counts have approximately Poisson noise,
this transformation implies that the variance of the output noise e(t)
is of order one. We also fixed the dynamics noise covariance so that
the prior equilibrium marginal variance of f was on the order 10, in
the units shown in the colorbars above; this is a fairly uninformative
prior. This left us with just one free hyperparameter tau (obtained
from the dynamics matrix, which was proportional to the identity, as
in the examples discussed in the paper). We tried tau=T/3 (left
panels) and tau=T/10 (right panels), to get a sense of of the
dependence of the estimate on tau. In all cases, we used a Gaussian
model for e(t), for simplicity. The place field was modeled as a
weighted sum of 400 equally-spaced, isotropic Gaussian bumps, with a
total resolution of 50x50=2500 pixels. Full forward-backward
smoothing required a few minutes on a laptop; the effective rank of
the posterior covariance minus the equilibrium covariance was on the
order of 20-40 throughout the experiment.
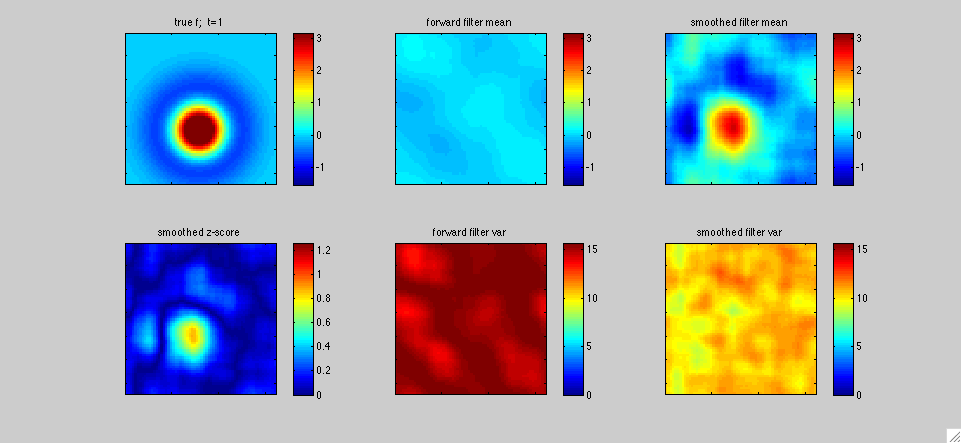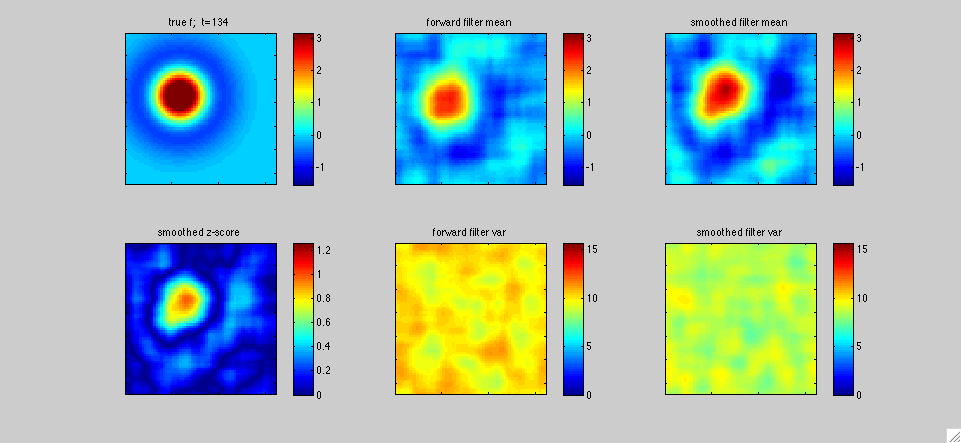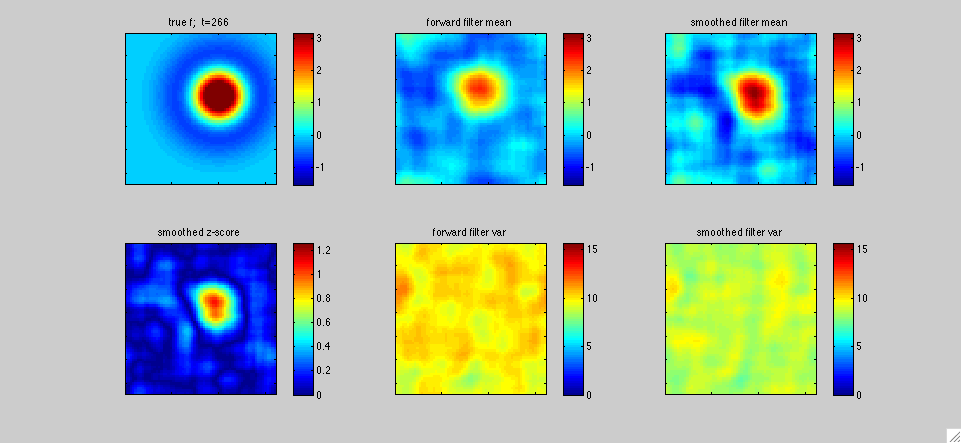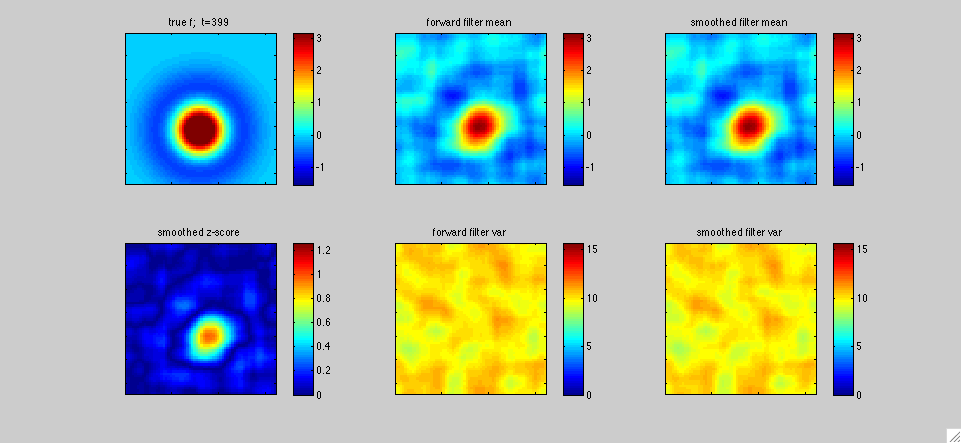
In
this second example, the left panel represents the receptive field
f[x,t] of a single simulated visual neuron as a function of the
two-dimensional visual spatial variable x and time t. The neuron's
observed output at time t, y(t), is represented as
y(t)=s(t)'f[.,t]+e(t), where e(t) is a random noise source, s(t) is
the visual stimulus at time t (not shown here; as in the
one-dimensional example discussed in the paper, we used spatiotemporal
white noise as the stimulus), and s(t)'f[.,t] denotes the dot product
of the stimulus with the receptive field at time t.
The top left panel shows the true receptive field f[.,t]; in this
case, the receptive field was chosen to be a difference of two spatial
isotropic Gaussian functions, and the position of these Gaussians was
varied sinusoidally in time. The middle panels display the fast
Kalman filter output at time t given the observations y(1),...,y(t)
resulting from the stimuli s(1),...,s(t), and the right panels display
the output of the Kalman smoother given the observed data over times
1,...T, with T=400 timesteps in this case. The lower left panel plots
a "z-score" statistic (the posterior mean in the upper right panel
divided by the square root of the posterior marginal variance in the
lower right panel), to give a sense of the signal-to-noise of the
output estimate.
The dimensionality of f[.,t] here was 64x64 pixels; tau = 100
timesteps; the power spectrum encoded in the prior covariance C0 fell
off with 1/w^2 (where w indexes the frequency of the Fourier basis),
as in the one-dimensional example discussed in the main text.
Liam
Paninski's home


